In August 2019, Piotr together with Andres Oberhauser organized a NSF-sponsored workshop “Progress and Prospects of Single-Molecule Force Spectroscopy in Biological and Chemical Sciences”.”.
Projects
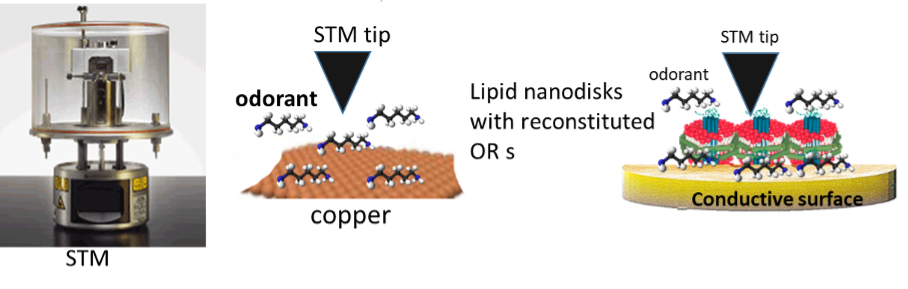
olfaction
Exploring the Quantum-Mechanical Basis of Odorant Detection by Olfactory Receptors
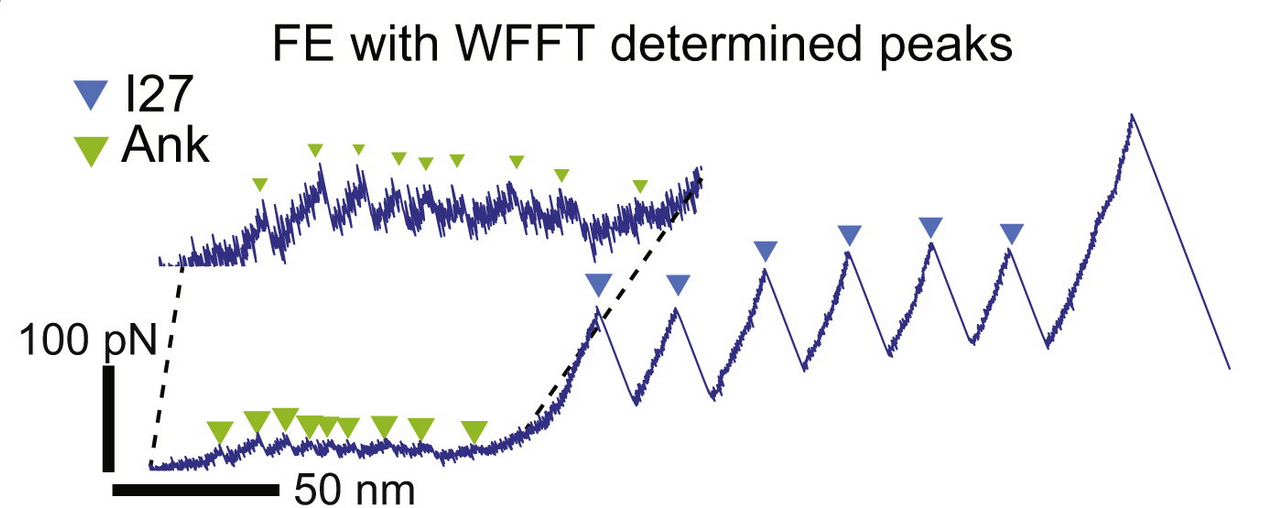
AFM design
Improvement of hardware and software for AFM

DNA Damage and Repair
Determining the mechanistic interactions between DNA damage and repair proteins
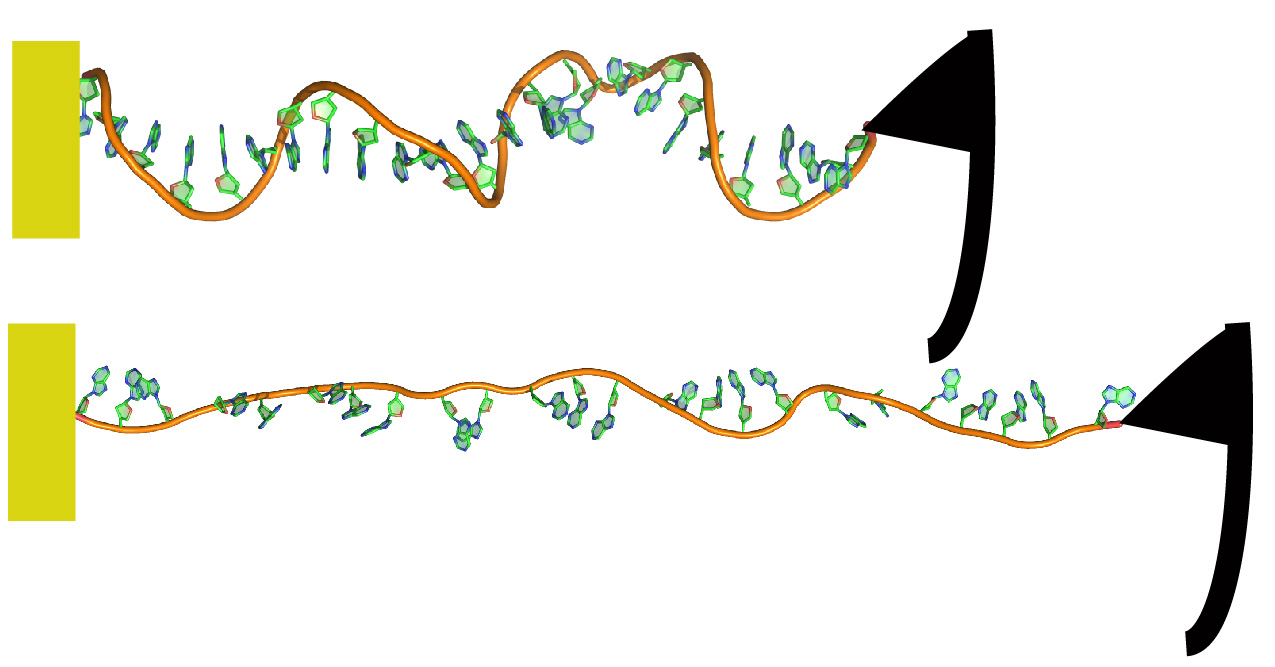
DNA Mechanics
Revealing the inherent mechanical properties of DNA
Mechanics of polysaccharides
AFM reveals the mechanical complexity of simple sugars
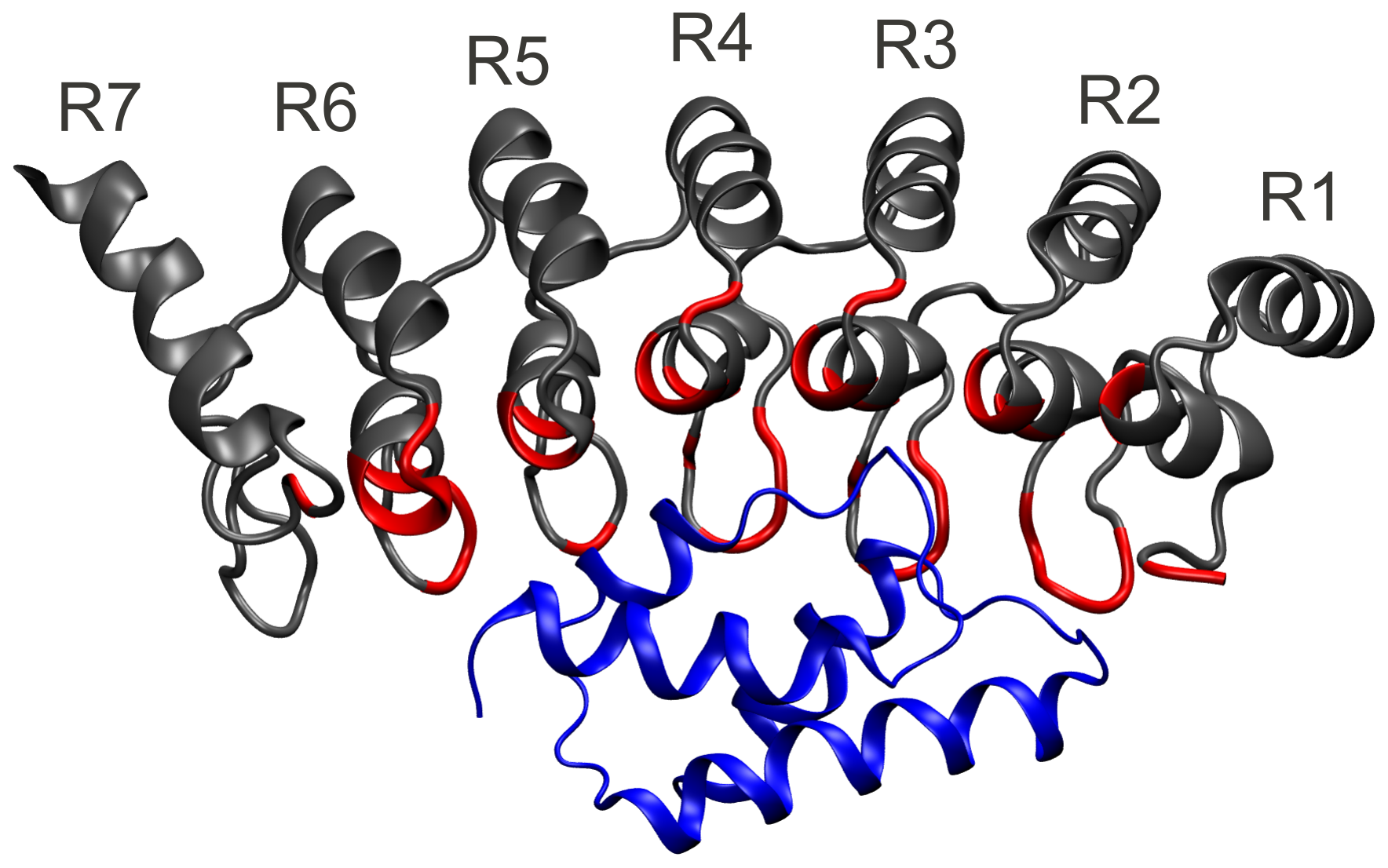
Nanomechanics of repeat proteins
Repeat proteins have many nice properties.

Protein folding
Repeat proteins have many nice properties.